AutoML with PyCaret for Hepatitis-C Prediction

Scientific Background
Hoffmann et al. (2018), in their paper "Using machine learning techniques to generate laboratory diagnostic pathways—a case study" challenged the use of expert rules in the interpretation of laboratory testing with machine learning models. The idea behind using machine learning is to provide an alternative to laboratory diagnostics that can offer state-of-art detection capabilities for certain conditions.
Fortunately, the data from this paper is available at the Machine Learning UCI Repository from UC Irvine. This dataset is also available at Kaggle as CSV, so I used the latter one.
The data was collected from 73 patients (52 males and 21 females), ages 19 to 75, with proven serological and histopathological diagnoses of hepatitis C. The data was labeled in several categories according to their hepatic activity index. This index was later used to transfer those groups into the machine learning classes.
The Data
Each entry in the dataset is described by ten biochemical tests, gender, and age. the following list details which test indicators were used in the data collection process:
Test Codes
- albumin (ALB)
- alkaline phosphatase (ALP)
- alanine amino-transferase (ALT)
- aspartate amino-transferase (AST)
- bilirubin (BIL)
- choline esterase (CHE)
- LDL (CHOL)
- creatinine (CREA)
- gamma glutamyl transpeptidase) (GGT)
- PROT
Additional attributes
- age
- sex
Response variable (y)
- Category (diagnosis) (values: '0=Blood Donor', '0s=suspect Blood Donor', '1=Hepatitis', '2=Fibrosis', '3=Cirrhosis')
Analysis with PyCaret
In the paper, the results using decision trees obtained the following results.
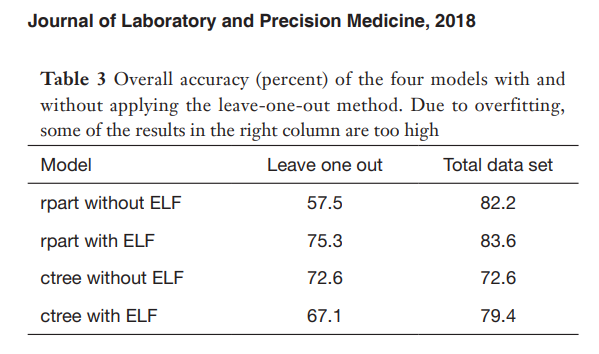
We are going to use the PyCaret AutoML capabilities to improve the accuracy over all the classes by using cross-validation.
Code
Imports
import pandas as pd
import numpy as np
from pycaret.classification import *
Load Data
data = pd.read_csv("HepatitisCdata.csv")
data = data.drop(labels='Unnamed: 0', axis=1)
data.head(10)
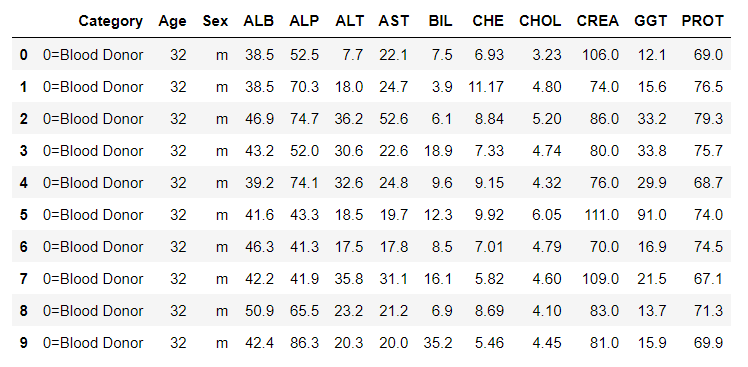
Data Preprocessing
# pd.Series(data["Category"], dtype="category")
# ['0=Blood Donor', '0s=suspect Blood Donor', '1=Hepatitis', '2=Fibrosis', '3=Cirrhosis']
data["Category"] = [0 if x == "0=Blood Donor" else x for x in data["Category"]]
data["Category"] = [1 if x == "0s=suspect Blood Donor" else x for x in data["Category"]]
data["Category"] = [2 if x == "1=Hepatitis" else x for x in data["Category"]]
data["Category"] = [3 if x == "2=Fibrosis" else x for x in data["Category"]]
data["Category"] = [4 if x == "3=Cirrhosis" else x for x in data["Category"]]
# pd.Series(data["Sex"], dtype="category")
# ['f', 'm']
data["Sex"] = [0 if x == "f" else 1 for x in data["Sex"]]
data.head(10)
The data looks much cleaner, and categorical columns were encoded accordingly.
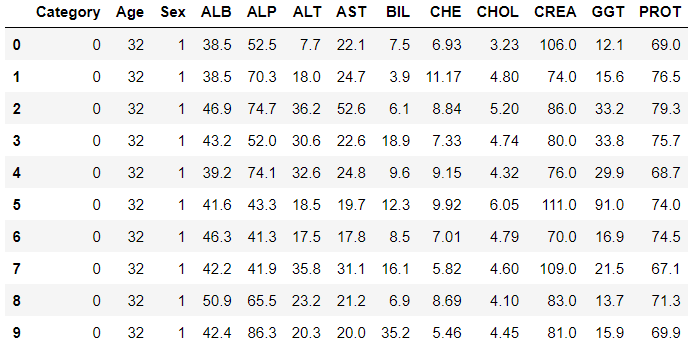
Magic with PyCaret
PyCaret with minimal coding will perform a benchmark of a suite of machine learning algorithms, will estimate metrics to determine which is the best model and will also calculate many visualizations such as AUC/ROC curves.
# pycaret setup
s = setup(data, target = "Category")
# model training and selection
best = compare_models()
This code snippet automatically created the following table of results, showing which algorithms performed better, and the metrics they aced are highlighted in yellow.
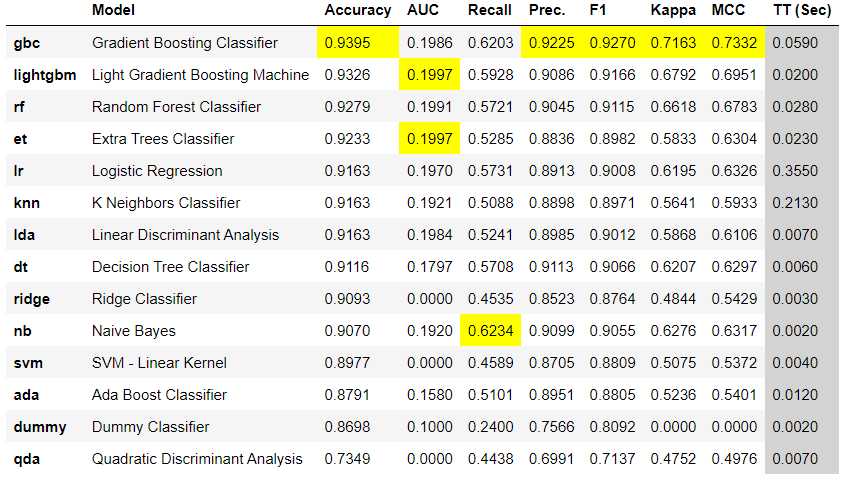
The clear winner here is the GBC Gradient Boosting Classifier algorithm.
Performance Metrics
PyCaret with a single line of code can provide a set of analyses that can help us identify how the mode is performing, accuracy, F1-Score, sensitivity analysis, and many more. Here are some interesting visualizations provided by PyCaret.
evaluate_model(best)
That's it! here are some analytics:
ROC/AUC Curve for GBC
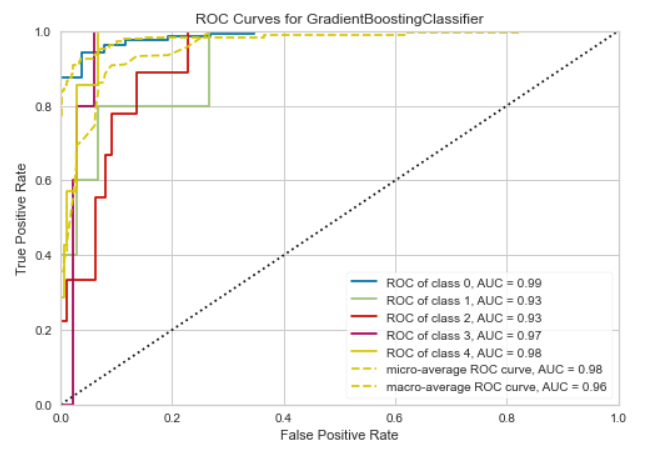
Confusion Matrix
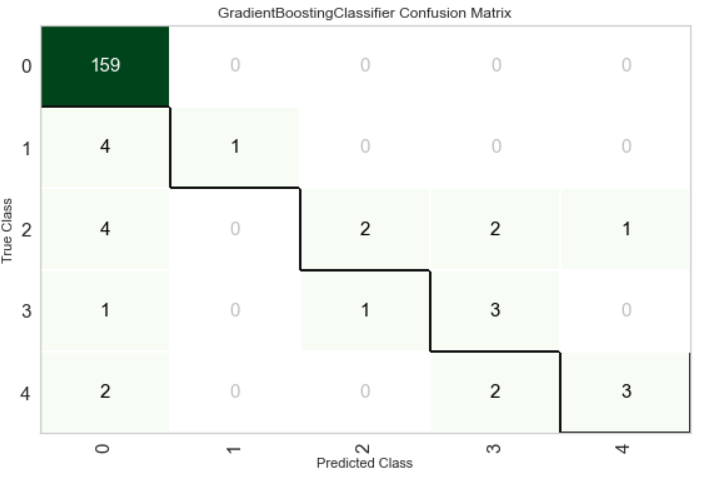
Precision-Recall Curve
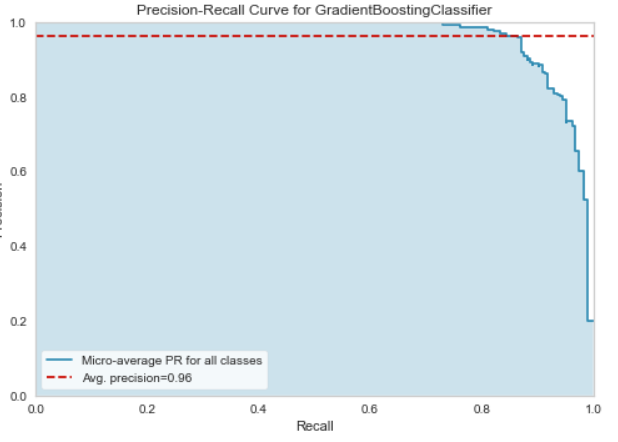
Classification Report
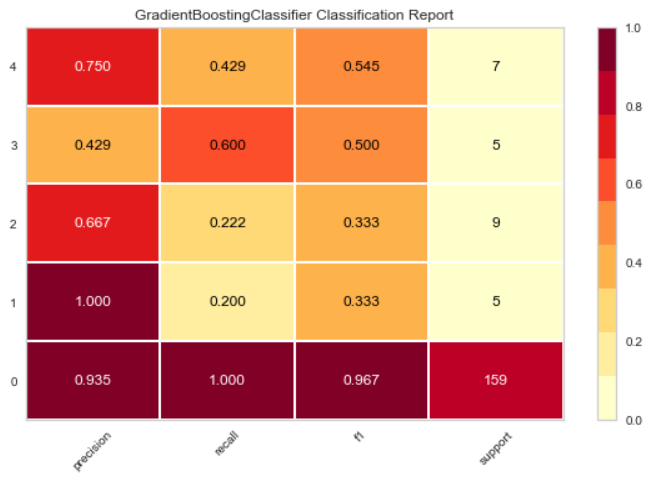
Prediction of New Patients
Before we start marking some predictions, PyCaret offers the save_model method to save the machine learning model as a pickle file. The following code shows how to save and load the model if you want later to use it in a web service.
Save Model
# save model to disk "hepatitis_model.pkl"
save_model(best, "hepatitis_model")
Load Model
from pycaret.classification import load_model
# load model
hepatitis_model = load_model('hepatitis_model')
Prediction of a new Patient
We are simulating a new patient, and we are creating a new pandas data frame with the information needed. the following snippet shows how to submit a new patient for inference.
Create the Data Frame
patient_data = pd.DataFrame()
patient_data = patient_data.append({
'Age': 32.00,
'Sex': 1.00,
'ALB': 44.30,
'ALP': 52.30,
'ALT': 21.70,
'AST': 22.40,
'BIL': 17.20,
'CHE': 4.15,
'CHOL': 3.57,
'CREA': 78.00,
'GGT': 24.10,
'PROT': 75.40
}, ignore_index=True)
Perform the Prediction
# perform prediction
prediction = predict_model(hepatitis_model, data = patient_data)
print('Patient was CLASSIFIED as:', classes[prediction.Label[0]])
The outcome of this prediction is: Patient was CLASSIFIED as: Blood Donor.
Summary
The PyCaret POC demonstrated a 0.9395 accuracy and 0.9270 F1-Score for predicting the classes in the diabetes dataset. This is way much better than the results shown by Hoffmann et al. Although this is a promising result, this code is still a POC and should not be used for production purposes until further evaluation by medical experts (I am, unfortunately, the wrong type of doctor here for medical and clinical approval!)
Download the Code
I uploaded the code a Gist Here!
if you liked this, remember to send some love back.






